One of the most promising and widely discussed use cases for AI in healthcare is using AI to diagnose different diseases. Due to increasing data availability and technological advancements, the scope of AI applications has expanded, covering sophisticated cases related to cancer detection and prediction.
However, the complexity of such solutions requires a cross-functional collaboration between medical and engineering experts. MobiDev has practical experience in training ML models for cancer diagnosis and supporting other digital healthcare workflows.
In this article, we’ll consider different AI use cases for cancer detection and share our expertise in this field.
TOP 4 AI Uses Cases for Cancer Detection
According to Global Market Insights, AI in oncology market size was valued at around $1.9 billion in 2023 and is estimated to reach over USD 17.9 billion by 2032. By providing insights, AI can support clinical decision-making. Let’s take a look at specific use cases.
1. Blood cell image analysis
Many types of cancer can be detected by analyzing blood cell images. However, the challenge lies in the sheer volume of blood cells, which makes manual analysis of all of them nearly impossible. Usually, doctors make decisions based on analysis of a limited number of cells, which is still very time-consuming.
The use of AI models can transform this process, enabling the analysis of all blood cells in automated mode. The models can generate reports for doctors highlighting key areas of concern which can significantly improve diagnostics.
2. AI for cancer detection and screening
Medical imaging approaches, such as computed tomography, magnetic resonance imaging, and positron emission tomography (PET) provide detailed insights into the structure of tissues and organs. Deep learning models, in turn, can be trained to analyze results, scans and images, and recognize conditions and patterns not identifiable by clinicians.
For example, the recent NCI-supported research confirms that AI improves breast cancer detection on mammography and helps predict the risks of breast cancers. Also, NCI researchers developed a deep-learning model for cervical and prostate cancer screening.
3. AI for tumor diagnosis
AI-powered tools, such as computer-aided detection (CAD) systems and machine learning (ML) algorithms, can analyze medical images to identify malignant tumors. For example, Support Vector Machines (SVM) and Artificial Neural Networks (ANN) demonstrate high accuracy in diagnosing prostate and breast cancers, often outperforming traditional diagnostic methods. While CAD systems effectively aid less experienced radiologists, they also reduce interobserver variability, accelerating diagnosis and improving outcomes.
Although DL models excel with large datasets, their application in clinical oncology faces challenges, such as data scarcity and interpretability. Radiomics, which involves extracting quantitative features from images, complements AI by enhancing tumor characterization and prognosis prediction.
4. AI for skin cancer detection
AI-powered skin cancer diagnostic devices are used to detect melanoma, basal cell carcinoma, and squamous cell carcinoma. For example, using spectroscopy technology, the handheld device can analyze skin lesions at a cellular and subcellular level, delivering quantitative, noninvasive results.
Combining AI with spectroscopy provides an efficient solution for evaluating lesions, enhancing both diagnostic precision and patient outcomes. With its ability to streamline care and improve detection, the device represents another prominent use case of AI in healthcare. Its adoption can help reduce missed diagnoses while increasing referral rates for suspicious cases.
In terms of implementation, the above-mentioned cases of AI cancer detection and prediction may rely on different approaches. Let’s take a look at them in more detail.
Current AI Approaches to Cancer Diagnosis
Machine learning and deep learning algorithms have demonstrated significant potential in identifying complex patterns within vast datasets, ranging from imaging to molecular biology. We can categorize all approaches as follows:
1. Machine learning algorithms
- Support Vector Machines (SVMs): Effective in molecular diagnostics and early cancer detection through feature selection in mass spectrometry data
- Random Forests: Widely used for predictive modeling, such as estimating cancer risk and identifying molecular biomarkers
- Decision Trees: Provide structured classification, often combined with ensemble methods to improve accuracy and reduce overfitting
- Gradient Boosting Algorithms (e.g., XGBoost): Excels in capturing nonlinear relationships and higher-order feature interactions within cancer datasets
2. Deep learning techniques
- Convolutional Neural Networks (CNNs): Central to image-based cancer detection, including tasks like lung nodule identification, skin lesion classification, and histopathological analysis
- Transformer Models: Emerging in clinical and genomic image processing for their ability to handle large datasets efficiently
- Long Short-Term Memory (LSTM): Applied to sequence data, including time-series imaging and longitudinal patient records
3. Microscopy-based AI
- Weakly Supervised Learning: Enables slide-level cancer prediction without detailed annotations, streamlining histopathological analysis
- Generative Adversarial Networks (GANs): Enhance imaging techniques, such as virtual staining and mitotic cell detection, improving diagnostic precision
4. Imaging-based AI
- Digital Breast Tomosynthesis (DBT): Incorporates deep learning classifiers to improve breast cancer detection rates and reduce false positives
- Low-Dose Computed Tomography (LDCT): Used in lung cancer screening, with AI algorithms identifying nodules and calculating malignancy risk
- Multi-Modality Integration: Combines imaging data (e.g., CT, MRI, PET) with molecular and clinical data for comprehensive cancer detection
5. Multimodal Approaches
- Integrated Omics Analysis: AI models combine genomics, transcriptomics, proteomics, and metabolomics data to uncover biomarkers and predict therapeutic responses
- Hybrid Models: Blend ML and DL techniques, such as deep support vector machines or ensemble learning, to improve accuracy in classification and survival prediction
Additionally, it is worth mentioning that Large Language Models (LLMs) are widely used in AI app development and represent a paradigm shift in natural language processing (NLP). These models apply advanced deep learning architectures to process and generate human-like text, facilitating tasks such as medical literature analysis, patient data summarization, and early cancer detection insights. LLMs can assist in extracting relevant data from unstructured EHRs, such as doctor notes and lab reports, to identify potential cancer cases or monitor disease progression.
MobiDev’s Experience in Training Models for Cancer Prediction
Modern histopathology involves multiple methods to analyze human cells, including tissue scans and anamnesis data. Today, healthcare organizations rely on Whole Slide Imaging or WSI to save a digital scan of the entire histology result. WSI scans help doctors analyze larger areas to detect malignant cells or other mutations in specific organ tissue.
However, the process of working with WSI is time-consuming, since such slides usually have large resolutions. Running through them manually is a time-consuming task. Given that this problem exists, the MobiDev team challenged themselves to search for an AI pathology solution capable of working with WSI and supporting digital healthcare workflows.
What is WSI?
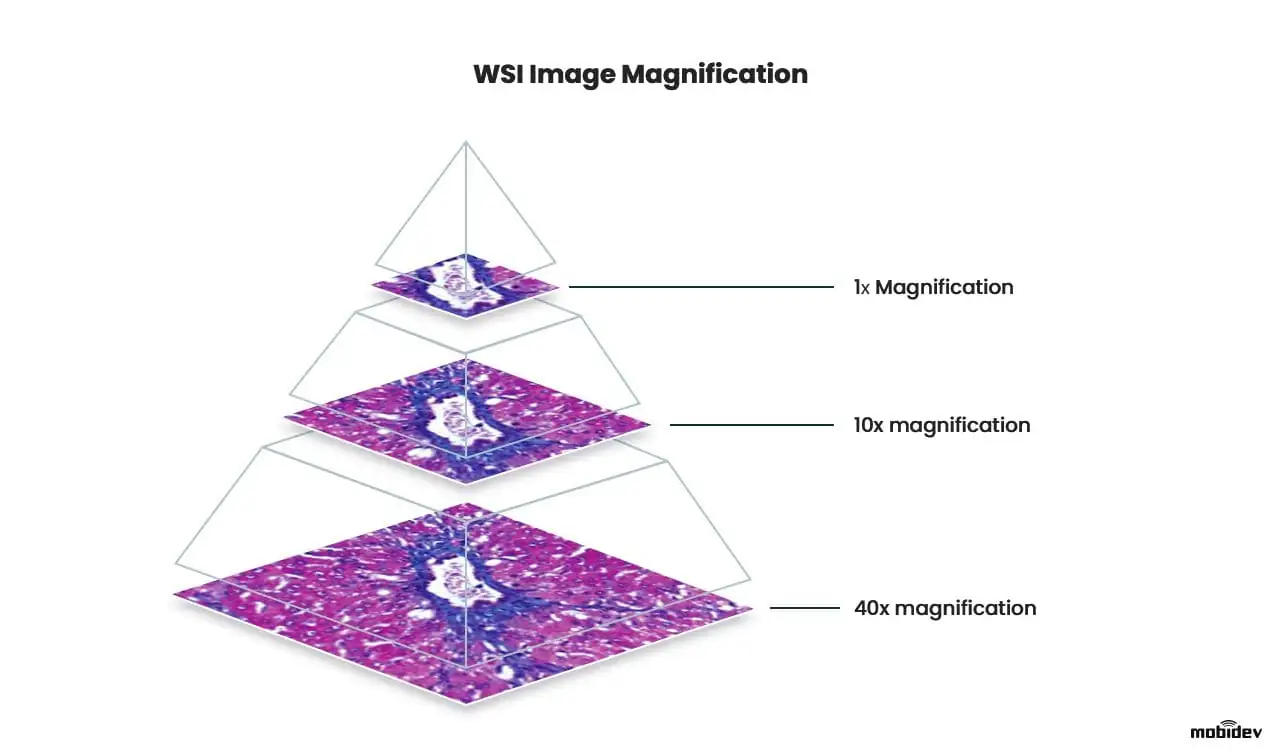
WSI image is a high-resolution, large-format scan of human tissue. WSI images are also called microscope slides, as they represent an enlarged image of cells and subcellular structure. Such slides are obtained by making multiple scans of different areas of a tissue sample and montaging them into one large image. As an output, we receive a digital scan that can be zoomed in up to 40 times without losing much detail.
WSI slide magnification
Compared to the traditional microscopic analysis of histology, WSI provides a number of advantages:
- Whole slide images usually come in giant resolutions like 100,000 x 100,000 pixels and bigger, which gives a lot of clarity when visually inspecting tissue areas.
- WSI is a digital scan that covers much larger areas all at once and is easier to zoom in/zoom out, compared to a microscopic view.
- The image can be sent between the healthcare experts as a digital copy. Additionally, WSI scans can’t deteriorate like specimens on a glass slide.
However, in terms of operability, WSI analysis is not so great due to the time and effort needed during the manual examination. In real-world practice, a doctor would sit before the image scanning by eye, zooming in and out for hours. In the case of manual examination, a whole slide should be examined to spot suspicious cell structures or mutations, because skipping some areas may result in medical error.
This is a general problem for healthcare workflows since long processes prevent quick diagnosis and efficient treatment, which are critical for patients with cancer and for histopathology as a whole. With the advances in digital pathology and fluctuations in healthcare regulatory requirements, the application of artificial intelligence became possible to use to process clinical data. As WSI is a gigapixel-sized image document that contains tons of unstructured data, the problem with manual examination can be addressed through machine learning.
Applying AI for WSI scan processing
Independent of the content in the image, WSI allows for the image to be broken down into features that are used for training machine learning models that run object detection, recognition, and classification tasks. These tasks are handled with computer vision that relies on optical sensors or digitally processes images to extract meaningful information.
However, since we don’t know where the potential tumor cells are, we can’t perform one essential ML task: data labeling. Labels are required to tell the model what these or those cells are for it to learn the difference between normal and damaged tissue. To solve this, convolutional neural networks, or CNNs, are used.
CNN requires little pre-processing of data input, as they are mainly used for visual data to find patterns autonomously. Applied to WSI scans, CNN is capable of extracting features out of weakly labeled, or completely unlabeled data. Extracted features then can be used to represent the whole slide in compact form, which can be helpful for image storing or grouping similar items.
However, minor pre-processing steps are still required, since it’s too CPU-demanding for the model to work with the whole slide. Due to the giant image size, the image is most likely to be sliced into equally-sized square images.
WSI processing through the CLAM approach
MobiDev approached the WSI processing system design through clustering. This is a constrained attention multiple instance learning (CLAM) approach. As a training dataset, the publicly available Clear Cell Renal Cell Carcinoma dataset was used.
To prepare data for training, images need to be cut into patches (equally sized square images), because the initial image size is too large. Also, the data used for the experiment was weakly labeled, meaning there were no labels for each patch of images.
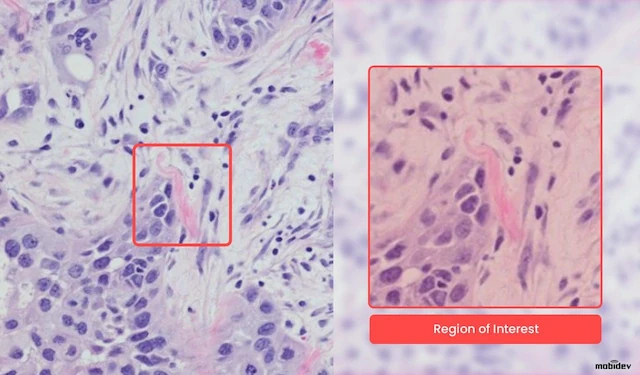
The task was to create a model capable of analyzing WSI scans and then highlight regions of interest, or in other words, suspicious cells, found in the tissue. As long as healthcare is based on critical decisions, the role of AI in pathology is to support the doctor with hints to speed up the diagnostic process.
However, the prediction should also be descriptive, as long as the user has to understand how the model derived this or that result. This is what’s called AI explainability as a feature of modern AI pathology systems.
CLAM is a data-efficient approach to working with weakly-labeled whole-slide imagery. The essence of it can be broken down into a number of steps:
- On the WSI, the tissue/cell sample is segmented from the background using computer vision algorithms.
- Within the segmented area, the tissue is split into equally sized square patches (usually 256px).
- Patches are encoded once by a pre-trained CNN into a descriptive feature representation. During training and inference, extracted patches in each WSI are passed to a CLAM model as feature vectors.
- For each class, the attention network ranks each region in the slide and assigns an attention score based on its relative importance to the slide-level diagnosis. Attention-pooling weighs patches by their respective attention scores and summarizes patch-level features into slide-level representations, which are used to make the final diagnostic prediction.
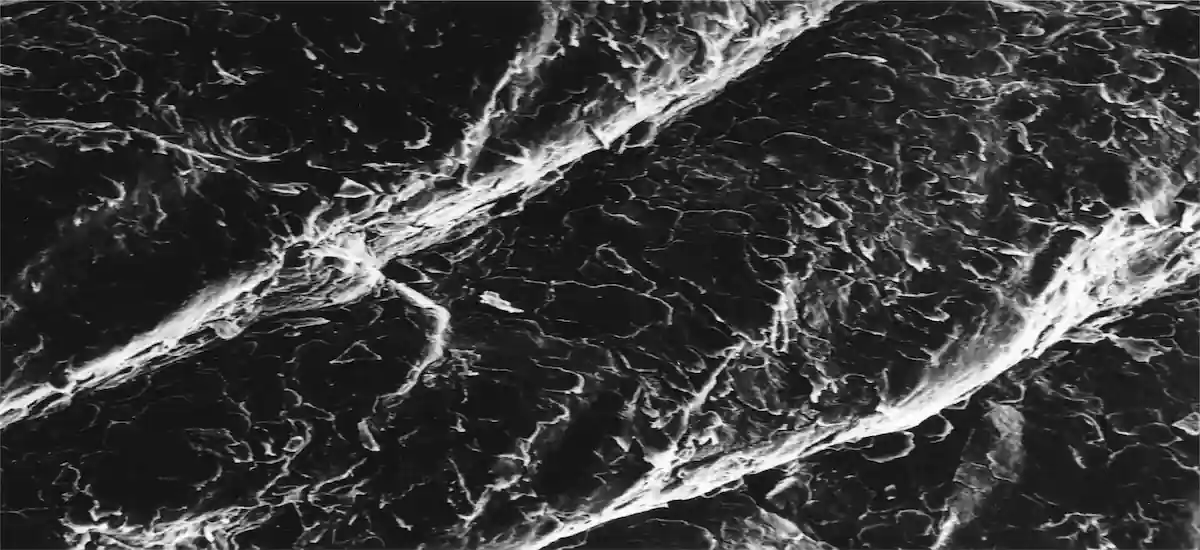
- The attention scores can be visualized as a heatmap to identify ROIs and interpret the important features.
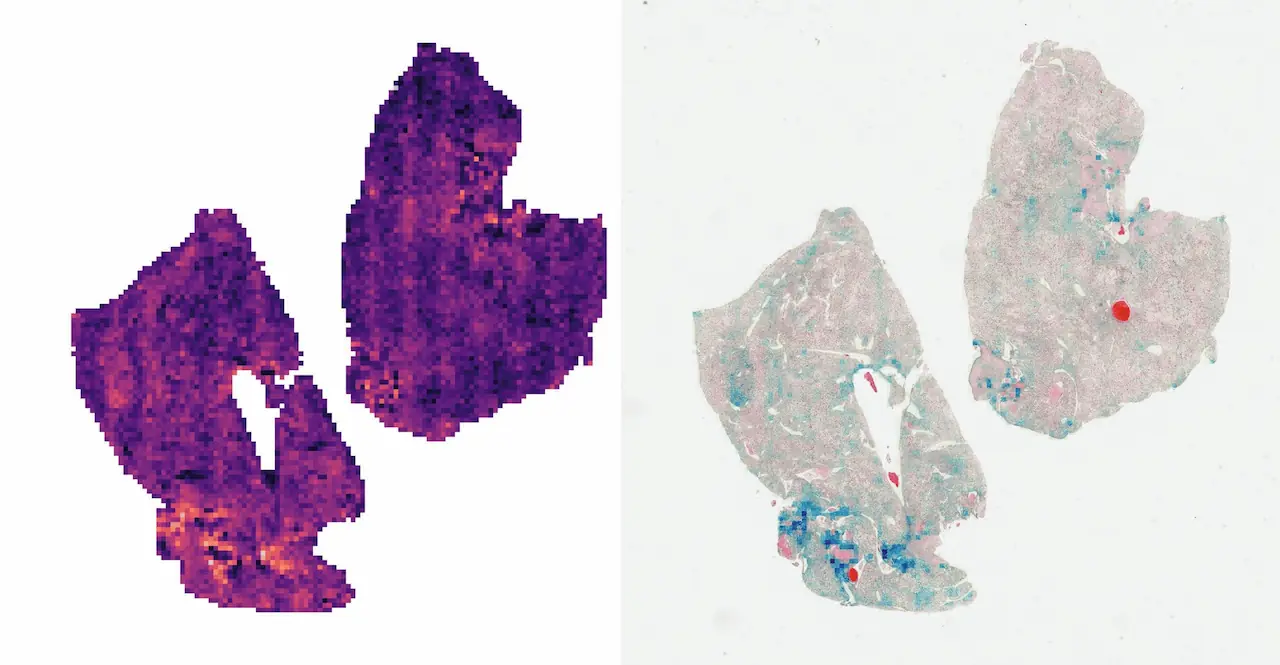
Left: direct visualization of the attention scores. Each pixel represents a 256px patch.
Right: attention scores blended onto the WSI. The colored regions represent the most attended parts of the sample.
As we can see, the CLAM’s attention network is quite interpretable. It outputs “importance scores” for each patch, so those can be visualized as a heatmap. If you take a closer look at the WSI on the right, you may see the affected cells colored blue, meaning the network determined they would affect the prediction the most.
Using neural networks to process weakly labeled WSI images provides a couple of advantages in this case. First of all, we can discover clinically relevant, but previously unrecognized morphological characteristics that have not been used in visual assessment by pathologists. Secondly, the model can recognize features that are beyond human perception, which can potentially provide more accurate diagnostic results and have a positive impact on surgical or therapeutic results.
Addressing the WSI complexity and interpretability
This approach addresses important WSI processing difficulties. For example, to handle the large size of the slide, we segment the actual tissue from the scan background, trimming unnecessary information. Then, we split the remaining ROI into tiles that are treated equally to fit them into the GPU.
Also, another difficulty is labeling. It takes hours for a pathologist to look through the slide and diagnose, let alone label it, i.e. identifying affected cells. On the contrary, CLAM is designed to work with this kind of weakly-labeled data, where only the slide-level labels are known (i.e. diagnosis.) CLAM itself determines which regions of the tissue affect the predicted diagnosis.
Other MobiDev’s AI Cancer Detection Researches
MobiDev has practical experience in more projects related to cancer detection tasks. For example, we worked on a solution to diagnose early signs of myelodysplastic syndrome by analyzing blood cells with AI models. The purpose of the developed system was to draw the attention of doctors to cases that require special attention and increase the probability of diagnosing the disease in the early stages.
Why Choose MobiDev as Your Engineering Ally for a Medical AI Product or Research
The MobiDev team has 15+ years of practical experience in healthcare app development services, building ML models for different needs, including cancer detection. We are committed to assisting research organizations with developing medical software and AI solutions that comply with HIPAA and security requirements.
Tap into our expertise if you need AI consulting services or AI development in the area of healthcare! We’ll be happy to help you with your mission.